Tutorials¶
The easiest way to get familiar with pertpy is to follow along with our tutorials. Many are also designed to work seamlessly in Google colab.
Note
For questions about the usage of pertpy use the scverse discourse.
Quick start¶
Guide RNA assignment
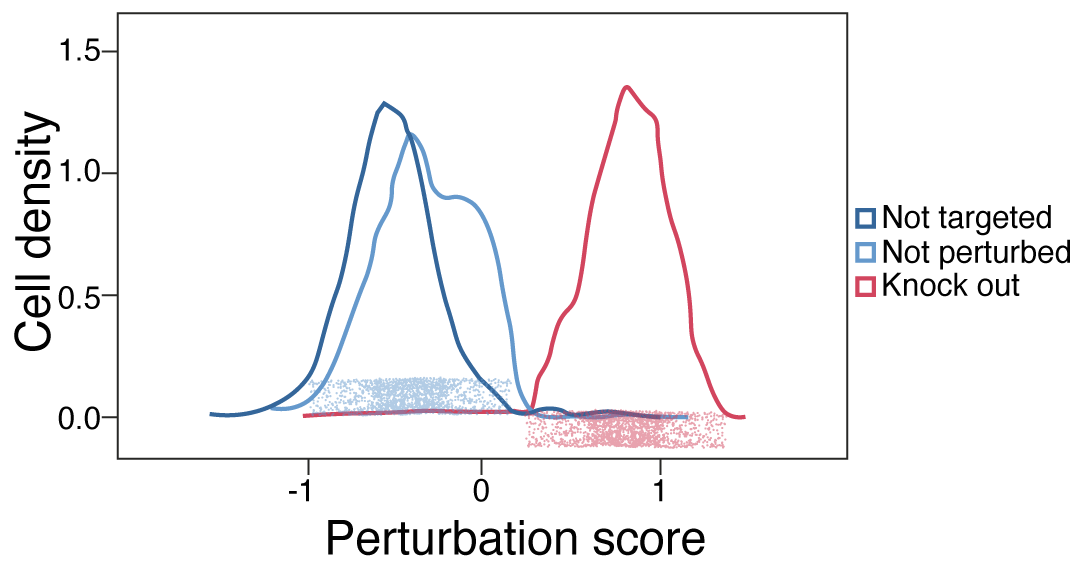
Mixscape - analysis of single-cell pooled CRSIPR screen
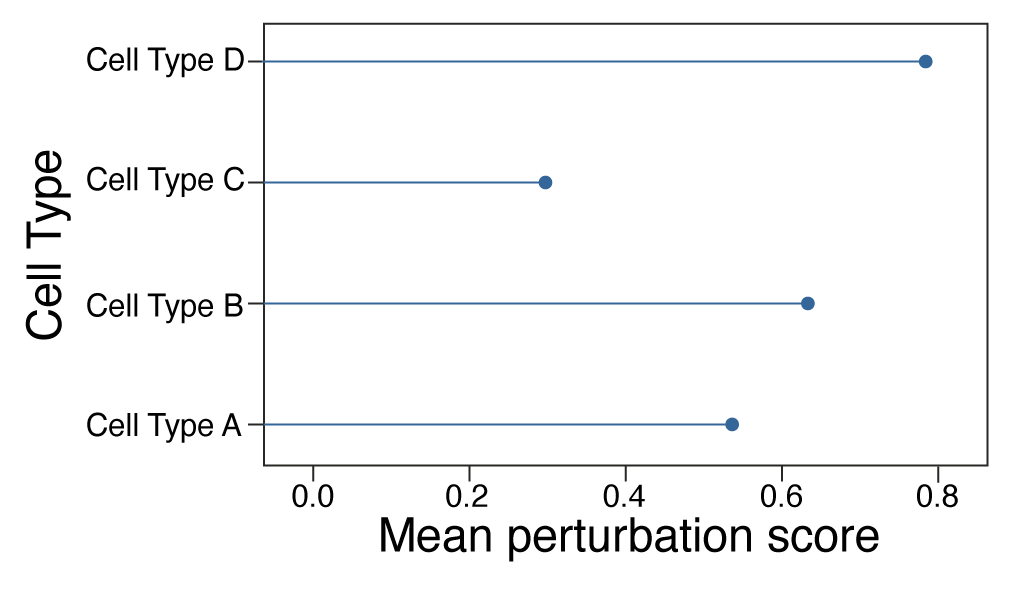
Augur - cell type prioritization prediction

scCODA - Compositional analysis of labeled single-cell data
scCODA - Modeling options and result analysis

tascCODA - Tree-aggregated compositional analysis
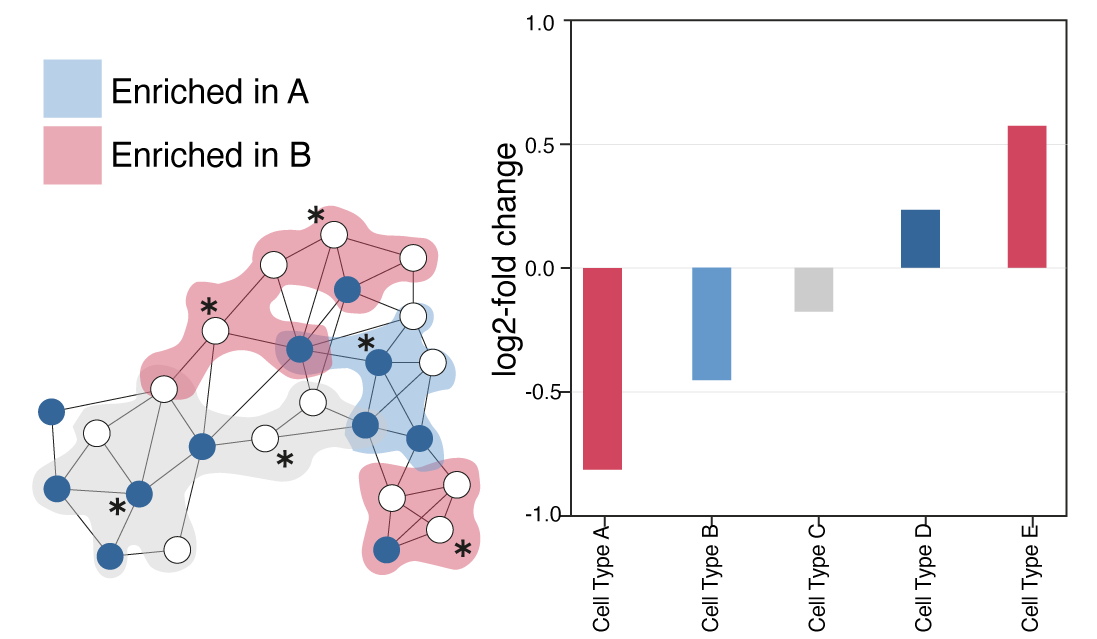
Milo - KNN based differential abundance analysis
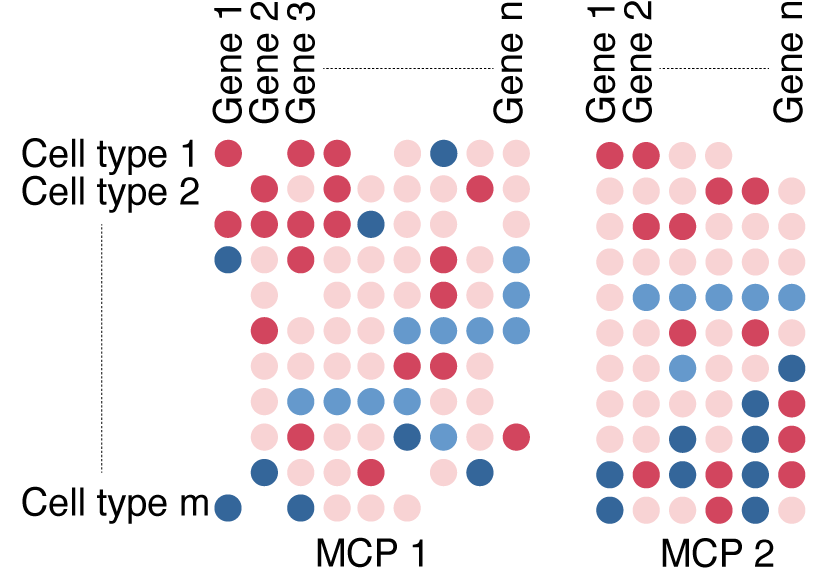
DIALOGUE - multi cellular programs

Enrichment

Distance metrics
Distance Tests
CINEMA-OT
scGen - Perturbation response prediction
Ontology mapping
Perturbation Space
Glossary¶
AnnData is short for Annotated Data and is the primary datastructure that pertpy uses. It is based on the principle of a single Numpy matrix X embraced by two Pandas Dataframes. All rows are called observations (in our case cells or similar) and the columns are known as variables (any feature such as e.g. genes or similar). For a more in depth introduction please read the AnnData paper.
For a more in depth introduction please read the Scanpy paper.